Time after time: calculating the autocorrelation function for uneven or grouped time series
I first learned how to check for autocorrelation via autocorrelation function (ACF) plots in R in a class on time series However, the examples we worked on were all single, long term time series with no missing values and no groups. I figured out later that calculating the ACF when the sampling through time is uneven or there are distinct time series for independent sample units takes a bit more thought. It’s easy to mistakenly ignore such structure, which then makes it difficult to determine what sort and how much autocorrelation may be present.
I first ran into this problem myself when I was analyzing data from a rotating panel study design. In the data I was working with, some units were sampled every year, some sampled every 3rd year, some every 9th year, etc. So sampling was annual, but not all sample units had observations from every year. In addition, the sample units were considered independent replicates of each time series, so any autocorrelation of concern would be within sample unit autocorrelation.
It took me some time to figure out how to check for overall residual autocorrelation from models I fit to these data. But I’m glad I took the time to do it; since then I’ve been able to share what I learned with numerous students when they were facing a similar situation with unevenness or grouped time series (or both!).
The kind of time series I’m talking about here are discrete time series, not continuous. Treating time as continuous involves a different approach, generally using “spatial” autocorrelation tools (which I may write about in some later post).
Table of Contents
- Simulate data with autocorrelation
- Fit model and extract residuals
- Problems with naively using acf()
- Calculate the maximum lag
- Order the dataset by time
- Pad the dataset with NA
- Plot autocorrelation function of appropriately-spaced residuals
- Add confidence interval to the ACF plot
- Just the code, please
Simulate data with autocorrelation
In my working example today I’ll use data that has a pattern to the unevenness, much like the data I had from the rotating panel design. The same approach applies, though, for evenly spaced data with groups or when some sampling events are missing because of unplanned events or logistical issues.
Autocorrelated noise can be simulated in R using the arima.sim() function. This thread on the R mailing list helped me figure out how to do this. I’m working with the default distribution of the innovations (i.e., errors) in arima.sim(), which is the normal distribution with a mean 0 and standard deviation 1.
I’ll use functions from purrr to do the looping and dplyr for data manipulation.
library(purrr) # v. 0.2.5
library(dplyr) # v. 0.7.5I’ll start the simulation by setting the seed. I mix things up in this post by using a seed of 64 instead of my go-to seed, 16. 😆
set.seed(64)I decided to simulate a 10-observation time series for 9 different sample units. Each sample unit is an independent time series so I do a loop via map_dfr() to simulate 9 separate datasets and then bind them together into one.
I’ll simulate observations of the response variable y and explanatory variable x for each time series and index time with an integer to represent the time of the observation (1-10). This time variable could be something like year or day or month or even depth in a soil core or height along a tree (since we can use time series tools for space in some situations). The key is that the unit of time is discrete and evenly spaced.
The response variable y will be constructed based on the relationship it has with the explanatory variable x along with autoregressive order 1 (AR(1)) errors. I set the lag 1 correlation to be 0.7.
The lag 1 correlation is the correlation between the set of observed values from time \(t\) with the values from time \(t\text{-}\mathit{1}\). In this example the lag 1 correlation for one sample unit is the correlation of the observed values at sampling times 2-10 with those at sampling times 1-9. The lag 2 correlation would be between the observations two sampling times apart (3-10 vs 1-8), etc. With 10 observations per group the largest lag possible is lag 9.
When you run the code to simulate the dataset below you will get warnings about not preserving the time series attribute from arima.sim(). I’ve suppressed the warnings in the output, but note these warnings aren’t a problem for what I’m doing here.
dat = map_dfr(1:9,
~tibble(unit = .x,
x = runif(10, 5, 10),
y = 1 + x + arima.sim(list(ar = .7), 10),
time = 1:10)
)Here’s the top 15 rows of this dataset.
head(dat, 15)# # A tibble: 15 x 4
# unit x y time
# <int> <dbl> <dbl> <int>
# 1 1 5.22 7.25 1
# 2 1 9.14 11.3 2
# 3 1 5.22 7.10 3
# 4 1 9.95 10.8 4
# 5 1 5.45 5.27 5
# 6 1 9.09 8.06 6
# 7 1 9.69 9.73 7
# 8 1 8.10 8.35 8
# 9 1 7.53 9.49 9
# 10 1 7.93 9.65 10
# 11 2 9.98 12.6 1
# 12 2 6.46 8.23 2
# 13 2 8.67 9.04 3
# 14 2 6.36 7.58 4
# 15 2 8.62 9.66 5The dataset I made has 9 sample units, all with the full time series of length 10. I want to have three units with samples taken only every fourth sampling time and three with samples on only the first and last sampling time. The last three units will have samples at every time.
I use filter() twice to remove rows from some of the sample units.
dat = dat %>%
filter(unit %in% 4:9 | time %in% c(1, 4, 7, 10) ) %>%
filter(!unit %in% 4:6 | time %in% c(1, 10) )Now you can see the time series is no longer even in every sample unit.
head(dat, 15)# # A tibble: 15 x 4
# unit x y time
# <int> <dbl> <dbl> <int>
# 1 1 5.22 7.25 1
# 2 1 9.95 10.8 4
# 3 1 9.69 9.73 7
# 4 1 7.93 9.65 10
# 5 2 9.98 12.6 1
# 6 2 6.36 7.58 4
# 7 2 6.51 7.91 7
# 8 2 6.79 7.34 10
# 9 3 6.71 8.52 1
# 10 3 8.64 10.1 4
# 11 3 9.00 7.74 7
# 12 3 9.49 9.91 10
# 13 4 5.20 3.25 1
# 14 4 9.52 11.3 10
# 15 5 9.26 10.8 1Fit model and extract residuals
I’m going to focus on checking for residual autocorrelation here, since that is what I do most often. This means checking for autocorrelation that is left over after accounting for other variables in the model. We often hope that other variables explain some of the autocorrelation. 🤞
I’ll fit the model of y vs x via the lm() function and extract the residuals to check for autocorrelation.
fit1 = lm(y ~ x, data = dat)I add the residuals to the dataset to keep things organized and to get ready for the next step.
dat$res = residuals(fit1)Problems with naively using acf()
If we hadn’t thought about our spacing issue in our grouped dataset, the next step would be to use the acf() function to check for any residual autocorrelation at various time lags. The acf() function calculates and plots the autocorrelation function of a vector of values. That sounds like what we would want to do; so why do I say this would be naive?
The acf() function expects values to be in order by time and assumes equal spacing in time. This means the acf() function considers any two observations that are next to each other in the dataset to be 1 lag apart even though it may have been three or more years since the last observation. If we calculate the autocorrelation function directly on the residuals as they are now we would ignore those instances where we didn’t sample for several years.
In addition, the acf() function doesn’t know we have independent groups. The last observation of one time series comes immediately before the first observation of another, so acf() will be consider these to be one lag apart even though they are unrelated.
If we want to work with the acf() function we’ll need to make sure that the dataset is spaced appropriately. To achieve this we can add empty rows to the dataset for any missing samples. In addition, we need to add rows between sample units so we don’t mistakenly treat them as if they were from the same time series.
Note that an alternative option for data like this, where we are assuming normally distributed errors, is to work with the nlme package. That package has an ACF() function that works on both gls and lme objects that will respect groupings.
Calculate the maximum lag
Since the units are considered to be independent of each other, the maximum lag to check for autocorrelation should be based on the maximum number of observations in a group. Below is one way to check on group size. Because I simulated these data I already know that the longest time series has 10 observations, but when working with real data this sort of check would be standard.
dat %>%
count(unit)# # A tibble: 9 x 2
# unit n
# <int> <int>
# 1 1 4
# 2 2 4
# 3 3 4
# 4 4 2
# 5 5 2
# 6 6 2
# 7 7 10
# 8 8 10
# 9 9 10Printing out the result wouldn’t be useful for a large number of groups, so here’s an alternative to look at just the maximum group size.
dat %>%
count(unit) %>%
filter(n == max(n) )# # A tibble: 3 x 2
# unit n
# <int> <int>
# 1 7 10
# 2 8 10
# 3 9 10Order the dataset by time
Since the acf() function is expecting a vector that is in order by time, always make sure things are in order prior to using acf(). I would use arrange() for this, putting things in order by group and then time within group.
dat = dat %>%
arrange(unit, time)Pad the dataset with NA
When I was working through this problem for the first time, I found these lecture notes that showed the basic idea of adding observations in order to get the spacing between groups right. In my own work I originally achieved this using some joins. Since then I’ve found switched to using the complete() function from package tidyr for this task.
library(tidyr) # v. 0.8.1You must have a variable representing the autocorrelation variable in the dataset for this approach to work. (This may seem obvious, but I’ve seen datasets that rely on the order of the dataset rather than having a specific time variable.) In this case that variable is time.
I group the dataset by unit prior to using complete() so every group has rows added to it. I define what values of time I want to be in the dataset within complete(). These values are based on 1., the sampling times present in the dataset and 2., the maximum group size. I need more rows between groups than the maximum lag I’m going to check for autocorrelation. The maximum lag I will explore is a lag 9 so I will add 10 extra rows between each sample unit in the dataset.
Since I’m working with an integer variable for time I can make the full sequence I want in each group via 1:20 (but also see tidyr::full_seq()). This means that I will add rows for times 1 through 20 for every sample unit in the dataset.
dat_expand = dat %>%
group_by(unit) %>%
complete(time = 1:20) Here is an example of what the first group looks like in the newly expanded dataset. It has rows added for the times that weren’t sampled along with 10 rows of NA at the end.
filter(dat_expand, unit == 1)# # A tibble: 20 x 5
# # Groups: unit [1]
# unit time x y res
# <int> <int> <dbl> <dbl> <dbl>
# 1 1 1 5.22 7.25 0.889
# 2 1 2 NA NA NA
# 3 1 3 NA NA NA
# 4 1 4 9.95 10.8 -0.746
# 5 1 5 NA NA NA
# 6 1 6 NA NA NA
# 7 1 7 9.69 9.73 -1.57
# 8 1 8 NA NA NA
# 9 1 9 NA NA NA
# 10 1 10 7.93 9.65 0.290
# 11 1 11 NA NA NA
# 12 1 12 NA NA NA
# 13 1 13 NA NA NA
# 14 1 14 NA NA NA
# 15 1 15 NA NA NA
# 16 1 16 NA NA NA
# 17 1 17 NA NA NA
# 18 1 18 NA NA NA
# 19 1 19 NA NA NA
# 20 1 20 NA NA NAPlot autocorrelation function of appropriately-spaced residuals
Now that things are spaced appropriately and in order by time, I can calculate and plot the residual autocorrelation function via acf(), using the residuals in the expanded dataset.
Note the use of na.action = na.pass, which is what makes this approach to work. The lag.max argument is used to set the maximum lag at which to calculate autocorrelation.
This is now a plot we can use to check for the presence and amount of autocorrelation. 🎉
acf(dat_expand$res, lag.max = 9, na.action = na.pass, ci = 0)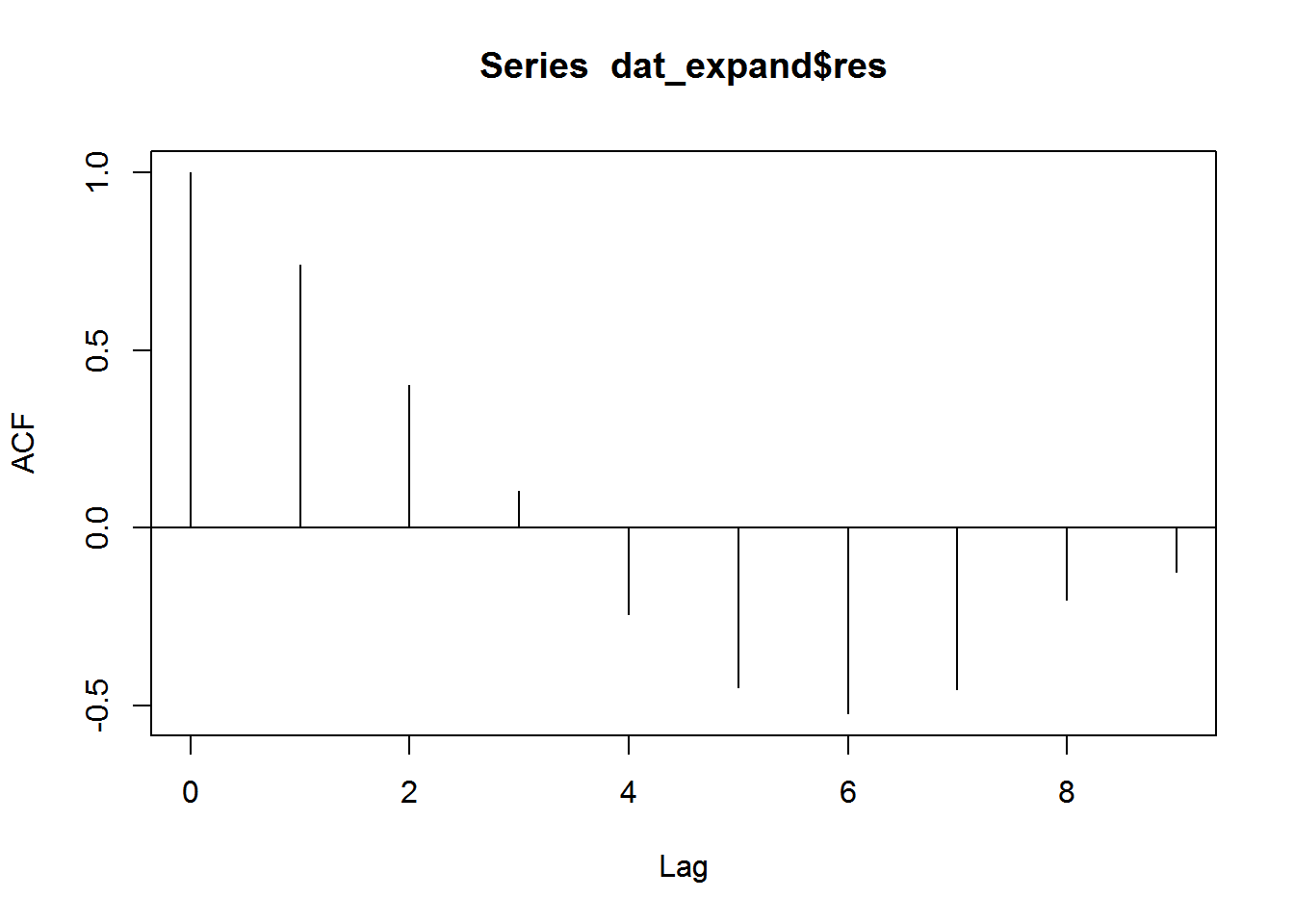
Add confidence interval to the ACF plot
I removed the confidence interval from the plot above with ci = 0. The confidence intervals calculated by acf() use the NA values as part of the sample size for each lag, so the confidence interval is too narrow.
You can calculate your own confidence interval based on the actual number of observations for each lag, but you have to calculate them yourself. I refer you again to those lecture notes for some ideas on how to do this. I’ll put code I’ve used for this in the past below as an example, but I won’t walk through it.
( nall = map_df(1:9,
~dat %>%
group_by(unit) %>%
arrange(unit, time) %>%
summarise(lag = list( diff(time, lag = .x ) ) )
) %>%
unnest(lag) %>%
group_by(lag) %>%
summarise(n = n() ) )# # A tibble: 9 x 2
# lag n
# <int> <int>
# 1 1 27
# 2 2 24
# 3 3 30
# 4 4 18
# 5 5 15
# 6 6 18
# 7 7 9
# 8 8 6
# 9 9 9Here’s the ACF plot with 95% CI added via lines().
acf(dat_expand$res, lag.max = 9, na.action = na.pass, ci = 0)
lines(1:9,-qnorm(1-.025)/sqrt(nall$n), lty = 2)
lines(1:9, qnorm(1-.025)/sqrt(nall$n), lty = 2)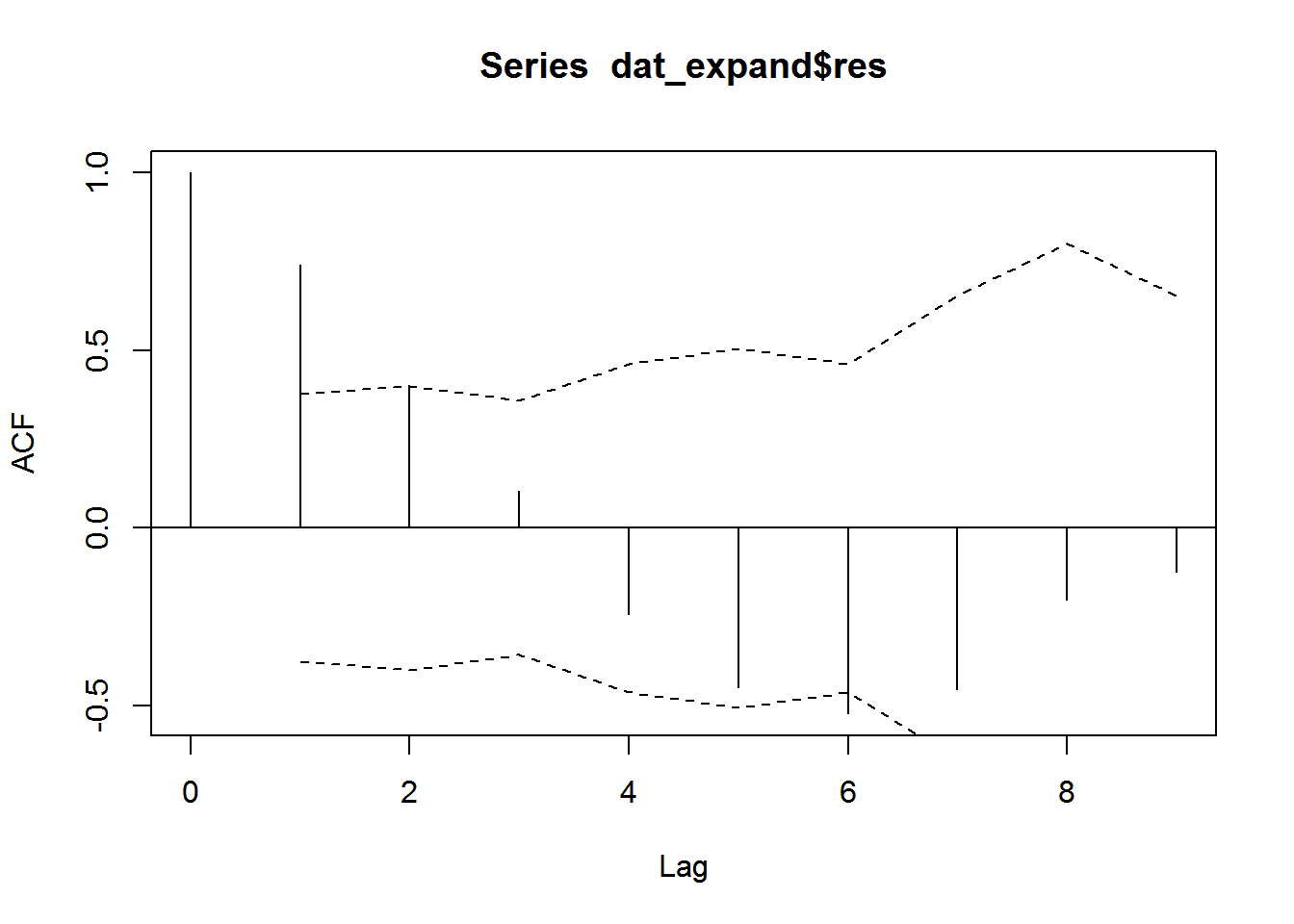
Just the code, please
Here’s the code without all the discussion. Copy and paste the code below or you can download an R script of uncommented code from here.
library(purrr) # v. 0.2.5
library(dplyr) # v. 0.7.5
set.seed(64)
dat = map_dfr(1:9,
~tibble(unit = .x,
x = runif(10, 5, 10),
y = 1 + x + arima.sim(list(ar = .7), 10),
time = 1:10)
)
head(dat, 15)
dat = dat %>%
filter(unit %in% 4:9 | time %in% c(1, 4, 7, 10) ) %>%
filter(!unit %in% 4:6 | time %in% c(1, 10) )
head(dat, 15)
fit1 = lm(y ~ x, data = dat)
dat$res = residuals(fit1)
dat %>%
count(unit)
dat %>%
count(unit) %>%
filter(n == max(n) )
dat = dat %>%
arrange(unit, time)
library(tidyr) # v. 0.8.1
dat_expand = dat %>%
group_by(unit) %>%
complete(time = 1:20)
filter(dat_expand, unit == 1)
acf(dat_expand$res, lag.max = 9, na.action = na.pass, ci = 0)
( nall = map_df(1:9,
~dat %>%
group_by(unit) %>%
arrange(unit, time) %>%
summarise(lag = list( diff(time, lag = .x ) ) )
) %>%
unnest(lag) %>%
group_by(lag) %>%
summarise(n = n() ) )
acf(dat_expand$res, lag.max = 9, na.action = na.pass, ci = 0)
lines(1:9,-qnorm(1-.025)/sqrt(nall$n), lty = 2)
lines(1:9, qnorm(1-.025)/sqrt(nall$n), lty = 2)